Purpose
Large-scale radiotherapy research increasingly relies on heterogeneous DICOM datasets containing complex cross-references across imaging and radiotherapy objects. Beyond ingestion, downstream preprocessing for analysis and machine learning requires consistent organization, validation, and transformation of these data at cohort scale. Efficient frameworks that unify ingestion, relationship resolution, visualization, and preprocessing remain limited. This work presents rosamllib, an open-source Python framework designed to support scalable radiotherapy DICOM processing, visualization, and data preparation for big-data research workflows.
Methods
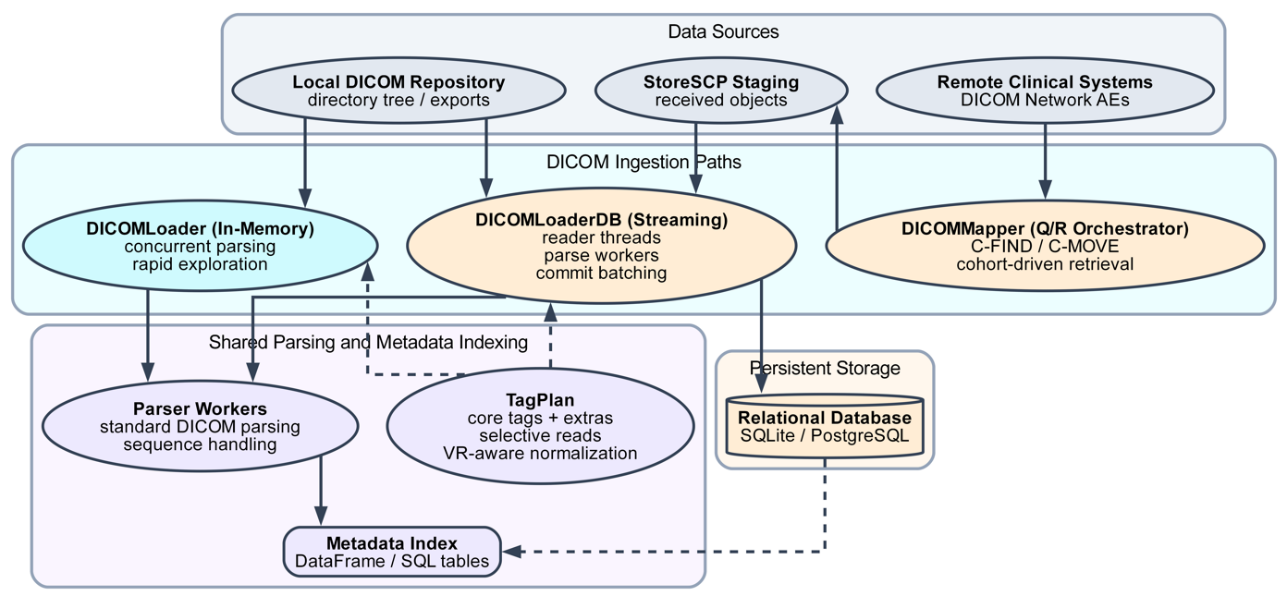
rosamllib provides complementary in-memory and database-backed ingestion pathways optimized for different data scales. DICOM objects are ingested from local filesystems or retrieved via standard DICOM query and retrieve operations and organized into a hierarchical, graph-based data model representing patients, studies, series, and instances. Cross-object relationships are explicitly resolved using DICOM identifiers and frame-of-reference associations. To support large datasets, the database-backed workflow employs a streaming producer–consumer architecture that decouples reading, parsing, and database writes, with selective tag extraction and value-representation–aware normalization. Built-in querying and visualization utilities enable cohort-level filtering and graphical inspection of series-to-series relationships. These capabilities support reproducible preprocessing workflows, including structure mask generation, dose alignment, data normalization, and export of model-ready datasets for deep learning training.
Results
The framework was applied to approximately one year of institutional radiotherapy DICOM data, comprising 1,869 patients, 12,406 studies, 349,246 series, and 6,370,243 instances. rosamllib enabled scalable ingestion, relationship validation, visualization, and automated preprocessing across large cohorts, supporting both exploratory analysis and machine learning–oriented data preparation.
Conclusion
rosamllib provides a scalable and extensible foundation for big-data radiotherapy research. By unifying ingestion, metadata indexing, relationship resolution, visualization, and preprocessing within a single framework, it enables efficient cohort-scale data preparation and supports downstream analytics and machine learning workflows in radiation oncology.
Innovation / Impact
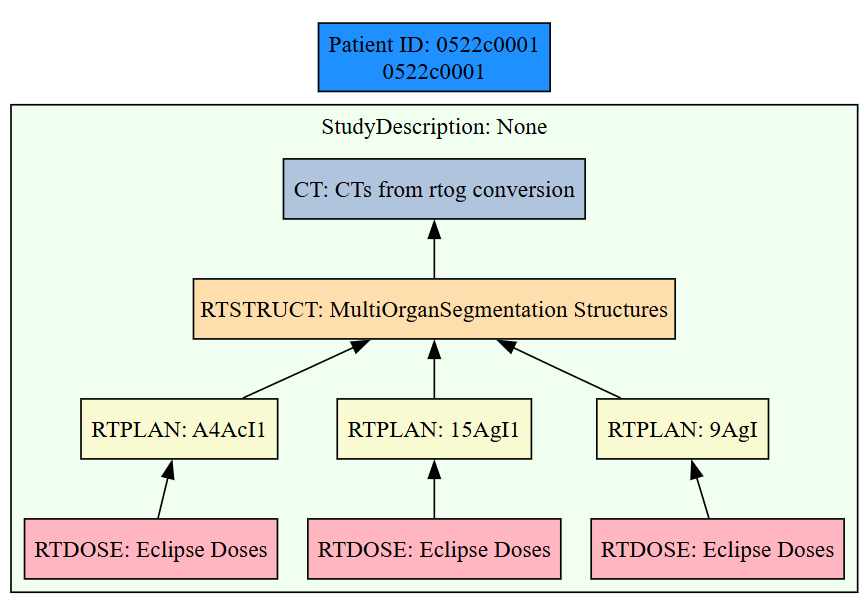
rosamllib introduces a unified framework for scalable radiotherapy DICOM processing that integrates ingestion, metadata indexing, and visualization within a single architecture. As illustrated in Figure 1, the framework supports both in-memory workflows for rapid data exploration and streaming, database-backed ingestion for cohort-scale processing, enabling efficient handling of large institutional datasets. A distinguishing feature of the framework is its explicit resolution and visualization of radiotherapy-specific inter-series relationships. As shown in Figure 2, rosamllib generates interpretable graphs linking imaging and treatment objects, allowing users to rapidly validate dataset structure and dependencies that are difficult to assess at scale. Together, these capabilities reduce the technical burden of large-scale radiotherapy data preparation and support reproducible preprocessing for downstream analytics and machine learning research.
Figures